![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SOX18_S5_R2_001.fastq |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 754700 |
| Filtered Sequences | 0 |
| Sequence length | 151 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality

![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores

![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content

![[WARN]](Icons/warning.png) Per base GC content
Per base GC content

![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution

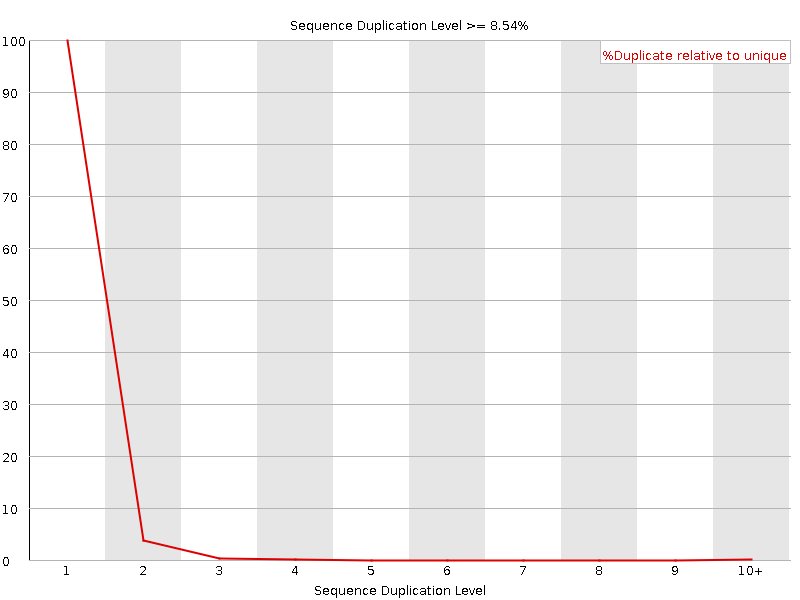
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG | 1283 | 0.17000132502981316 | No Hit |
| GATCTGGTTTTATTACATTCTGAATTGGACGTTGAAAATGAGCTTATCTC | 1200 | 0.15900357758049558 | No Hit |
| GATCCAGCCATAAAATGCATCATTCTTTTTTGTTTTAGACAACATTTCAT | 1186 | 0.15714853584205646 | No Hit |
| GATCCAGCTATAAAATGCATCATTCTTTTTTGTTTTAGACAACATTTCAT | 975 | 0.12919040678415264 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
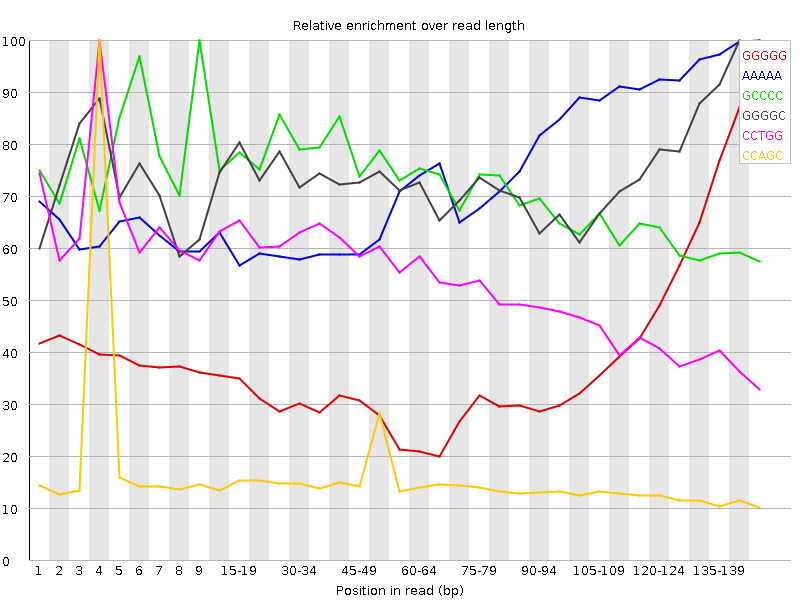
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 685800 | 13.205257 | 34.17288 | 145-147 |
| AAAAA | 804175 | 3.1815584 | 4.2258124 | 145-147 |
| GCCCC | 124575 | 3.0646746 | 4.317182 | 9 |
| GGGGC | 149610 | 3.0627463 | 4.1093903 | 140-144 |
| CCTGG | 169070 | 2.788919 | 5.3300524 | 4 |
| CCAGC | 152985 | 2.579582 | 17.694656 | 4 |
| ATCTC | 253950 | 2.3135746 | 8.164064 | 2 |
| GATCT | 267280 | 2.2903438 | 32.177734 | 1 |
| TCCAG | 163635 | 1.9669508 | 14.879388 | 3 |
| CAGCC | 112450 | 1.8960941 | 11.168093 | 5 |
| TTTTA | 389410 | 1.8029183 | 5.7797637 | 8 |
| CAGCT | 143380 | 1.7234786 | 5.331216 | 5 |
| GTTTT | 269195 | 1.7103617 | 6.0429125 | 7 |
| GATCG | 134605 | 1.5218697 | 11.687004 | 1 |
| TTTAT | 328100 | 1.5190609 | 5.429038 | 9 |
| ATCAT | 232680 | 1.4529185 | 5.4717464 | 2 |
| GATCA | 174880 | 1.4408025 | 25.185228 | 1 |
| ATCTG | 159405 | 1.3659543 | 14.728083 | 2 |
| AGCCA | 117760 | 1.3609595 | 7.774965 | 6 |
| TCTGG | 112785 | 1.3262872 | 9.252169 | 3 |
| ATCCA | 135345 | 1.1855164 | 15.306009 | 2 |
| CTGGT | 98245 | 1.155305 | 7.68574 | 4 |
| GGTTT | 135305 | 1.1342711 | 6.7479076 | 6 |
| ATCAG | 137640 | 1.1339893 | 7.6555934 | 2 |
| TGGTT | 124535 | 1.0439855 | 5.829416 | 5 |
| ATCAC | 118715 | 1.0398506 | 5.8983192 | 2 |
| GCCAT | 83695 | 1.0060437 | 5.1291924 | 7 |
| ATCAA | 154380 | 0.926837 | 6.1435447 | 2 |
| GATCC | 75700 | 0.9099409 | 36.31172 | 1 |
| ATCTT | 130470 | 0.84734994 | 5.8343277 | 2 |
| ATCCC | 63955 | 0.8173214 | 6.435413 | 2 |
| ATCCT | 84205 | 0.7671374 | 6.436149 | 2 |
| GATTA | 107250 | 0.62990993 | 5.9414773 | 1 |